Welcome to Toxic Protein Predictor for Plantae Web Server
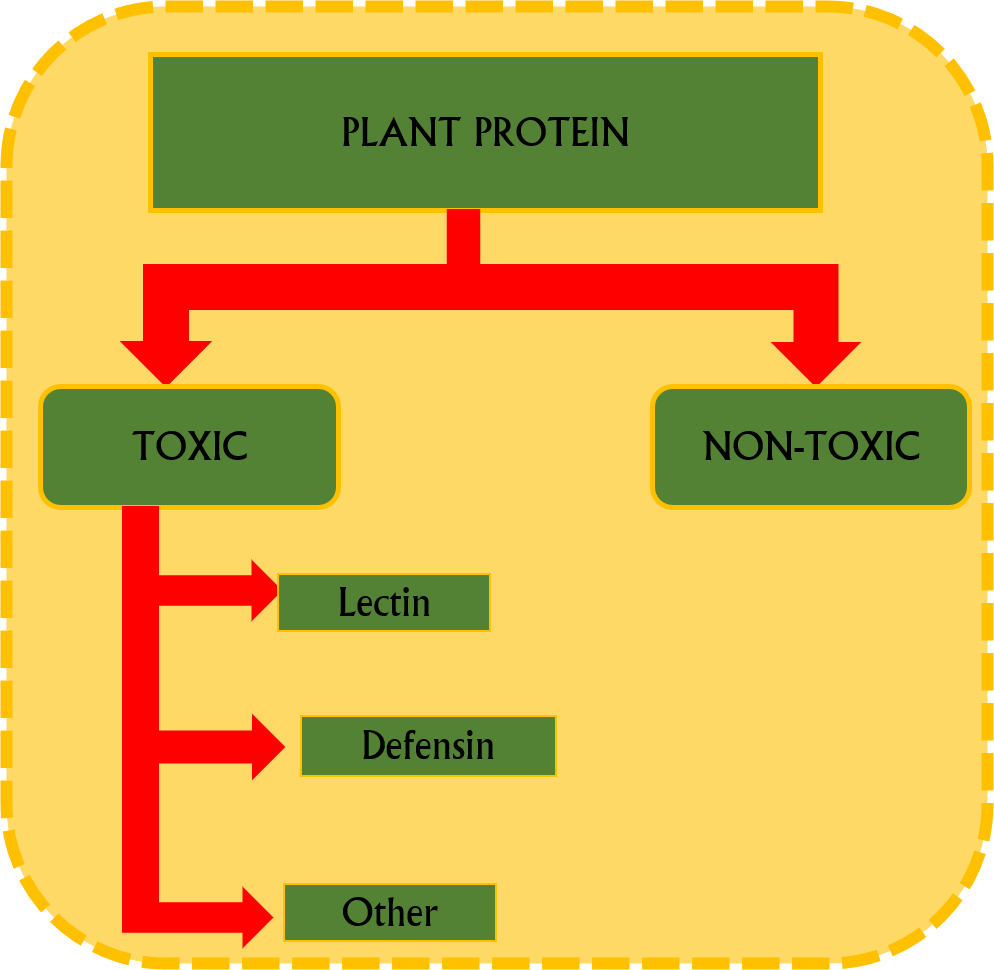
This server allows you to predict protein toxicity and run the full toxcity pipeline.Plants have evolved complex and highly regulated defense mechanisms to protect themselves against pathogenic invasion. One such mechanism involves the biosynthesis and secretion of toxic proteins that inhibit pathogen proliferation or induce pathogen cell death. These defense-related proteins include lectins, defensins, protease inhibitors, and chitinases, which interfere with vital metabolic pathways such as protein synthesis, cell wall formation, and membrane integrity in pathogens. Additionally, certain toxic proteins promote the generation of reactive oxygen species (ROS), thereby establishing an oxidative environment detrimental to microbial survival. Elucidating these molecular defense responses is crucial for advancing strategies in disease-resistant crop improvement. Recently, machine learning (ML) approaches have emerged as powerful computational tools for predicting toxic versus non-toxic plant proteins using sequence-derived features. Protein sequences can be transformed into numerical descriptors based on amino acid composition, physicochemical properties, and advanced embedding representations. Feature selection techniques further refine model performance by minimizing redundancy and enhancing discriminatory power. Integration of ML with large-scale proteomic and genomic datasets facilitates high-throughput screening of potential plant toxins, accelerating discovery pipelines, and enabling targeted experimental validation for improved crop resilience and safety assessment.